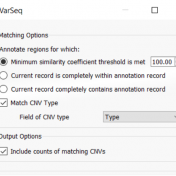
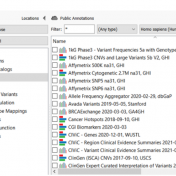
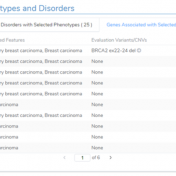
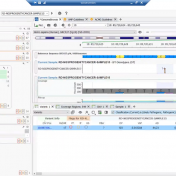
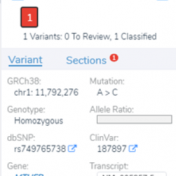
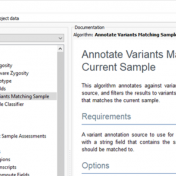
In continuation of our blog posts focusing on new features of VarSeq v2.2.2, here we will discuss the Latest Sample Assessment algorithm for both single nucleotide variants (SNVs) and copy number variants (CNVS). This algorithm annotates the variants of the project with the latest assessment from your variant catalog, which will show the history of interpretations made for the variants… Read more »
Annotating genomic variants is a very complex process but perhaps the most important part of next-generation sequencing variant analysis. Here at Golden Helix, we recognize the importance and value of having the most up-to-date sources available and curating new annotation sources as they become available for variant analysis. Golden Helix has curated over 100 annotation sources for human variant analysis… Read more »
Our recent release of VarSeq 2.2.2 comes with a long list of upgrades and new features. In this blog post, we will demonstrate how defining sample phenotypes are available in VSClinical. One noticeable change is the ACMG guideline variant evaluation in VSClinical. Not only has this interface added CNV guideline evaluation, simplified the reporting process with embedded Microsoft Word and… Read more »
Didn’t catch the webcast live? No worries! We cover ‘VSClinical: A Complete Clinical Workflow Solution’ Q&A’s in this blog post. The webcast, ‘VSClinical: A Complete Clinical Workflow Solution’ demonstrated how solutions provided by Golden Helix can be implemented to cover all requirements of a clinical workspace. Specifically, this webcast focused on a detailed workflow from a bioinformatician, geneticist, and lab… Read more »
Golden Helix continues to innovate and grow, and due to this, we have been garnering attention and praise. We are thrilled to share with you some of the accolades and interest from numerous publications from the last six months. Talking points ranged from diagnosing and tracking Covid-19 infections leveraging NGS to new workflows, interpreting and reporting on copy number variants… Read more »
Golden Helix offers a market-leading bioinformatics solution that allows users to evaluate next-generation sequencing variants according to the ACMG and now ACGS guidelines. The ACMG guidelines were created in 2015 and are widely accepted as best practice for the interpretation of sequencing variants throughout the United States (Richard et al 2015). Very similar are the ACGS guidelines that were developed… Read more »
Happy New Year! I hope you had an opportunity to relax over the holidays with your family and friends. It’s time to start talking about our plans for the New Year, but first, please allow me to review a few highlights from 2020 that helped build a foundation for our future. Growth: Golden Helix was named to the 2020 Inc… Read more »
Have you seen us in The Journal of Precision Medicine? Last week, our team released VarSeq 2.2.2 loaded with a number of updates and VSClinical’s highly-anticipated ACMG-CNV Guideline workflow! We have spent the past several months sharing webcasts and blogs on this new capability. We are honored to also have this new solution recognized in The Journal of Precision Medicine… Read more »
Reading scientific articles that our customers have recently published is one of my favorite things here at Golden Helix. It is fascinating to learn about the research and to see the various ways our software gets put to the test.Since we are rolling out our most extensive VarSeq update yet, I thought it would be a great time to look… Read more »
Our upcoming release of VarSeq is one of the largest we’ve ever had with our software! It comes with an extensive list of polishes and new features like our recently mentioned ACMG CNV classifier and a redesigned reporting interface with updated templates. Additionally, this new release is also paired with some major upgrades to our list of new and supported… Read more »
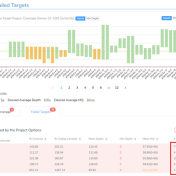
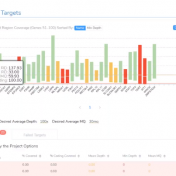
Curious about how coverage statistics can be used in conjunction with VarSeq? Evaluating the coverage over target regions or whole genomes is essential whether you are working with variant or CNV analysis. VarSeq has had the capability to compute sample level coverage statistics for some time now, but in the 2.2.2 release of VarSeq, there are some new features that… Read more »
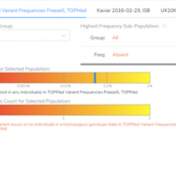
Our previous webcast demonstrated some of the new functionalities of VSClinical, including the ability to add ACMG frequency sources for the ACMG BA1, BS1, and PM2 criteria. This new feature was spurred by the feedback from our users, which requested supporting frequency tracks other than gnomAD Exomes and 1kG Phase3. Now, users can implement population catalogs to VSClinical such as… Read more »
Webcast Recap In the recent webcast “Exploring New Features and Clinical Reports in the ACMG Guideline Workflow”, Gabe and I took viewers through an evaluation with CNVs and SNVs according to the ACMG Guidelines where we generated and customized a clinical report. Along the way, we highlighted many new features that will soon be available in the upcoming VarSeq release…. Read more »
In this blog post, I will be analyzing a loss-of-function splice variant in MTHFR using VarSeq. In the search for clinically relevant variants contributing to rare disorders, efficient filtering strategies are an important step in eliminating disinteresting variants. However, any applied filters must also ensure no interesting variants inadvertently get filtered out. Golden Helix provides the tools to complete this… Read more »
In our previous webcast, Evaluating CNVs with VSClinical’s New ACMG Guidelines, we focused on a CNV deletion (12:27715515-29628122×1) in which the patient had a known disorder called Brachydactyly type E. The CNV was isolated using our VS-CNV caller and applied to the ACMG CNV guidelines using the intuitive steps of VSClinical. If you missed the webcast, you can watch the… Read more »
Those of you who have been attending our recent webcasts have learned about our upcoming VarSeq release. A part of that release will be an additional algorithm that will annotate variants matching the current sample. If you are not familiar with these webcasts, here are several on-demand webcasts I recommend to get you familiar with these new features: Evaluating Copy… Read more »
In the webcast, Evaluation of Copy Number Variants with VSClinical’s New ACMG Guideline Workflow, we discussed how VSClinical implements Section 4 of the ACMG guidelines. Specifically, we focused on integrating literature and publications to assess the pathogenicity of a CNV event when there was a lack of dosage sensitivity information. One of the primary pieces of evidence for evaluating genes… Read more »
Our team is regularly updating and curating annotations, most recently the BRCA Exchange. Breast cancer is known to occur in approximately 10% of the female population and if there is a damaging mutation in the breast cancer gene (BRCA), that rate increases to 65%. Although genetic testing can identify mutations in BRCA, a tumor suppressor gene, it has been difficult… Read more »
In lieu of conferences making the absolutely necessary decision to cancel this year’s conference, we created our Virtual 2020 Conference Booth! Even though it will never compare to the real deal, we hope you enjoy this blog post containing a few events we had planned this year. To start, let us paint the picture with some of our favorite conference… Read more »
Golden Helix is excited to release an upcoming VSClinical feature that allows users to analyze next-generation sequencing (NGS) CNV event reporting with ACMG guidelines. This feature will be the first in the NGS workspace to allow this capability and if you are curious about the functionalities you can get a sneak peek by looking at some of our most recent… Read more »